Exploring Connectivity of the Brain's White Matter
with Dynamic Queries
Stanford University
Stanford University
Stanford University
Stanford University
Stanford University
To appear in Transactions on Visualization and Computer Graphics in 2005 (This is an extended version of our IEEE Visualization 2004 paper.)
Paper
Video (interface demonstration)
Software
Abstract
Diffusion Tensor Imaging (DTI) is a magnetic
resonance imaging method that can be used to measure local
information about the structure of white matter within the
human brain. Combining DTI data with the computational
methods of MR tractography, neuroscientists can estimate the
locations and sizes of nerve bundles (white matter pathways)
that course through the human brain. Neuroscientists have used
visualization techniques to better understand tractography data,
but they often struggle with the abundance and complexity of the
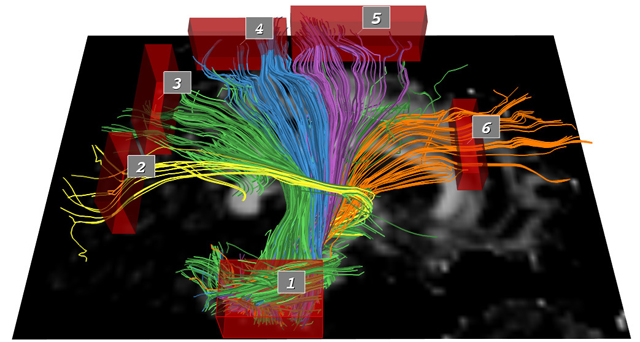
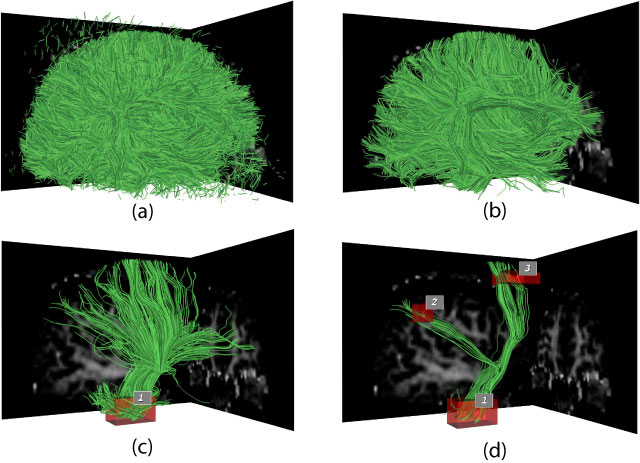
pathways. In this paper, we describe a novel set of interaction
techniques that make it easier to explore and interpret such
pathways. Specifically, our application allows neuroscientists
to place and interactively manipulate box- or ellipsoid-shaped
regions to selectively display pathways that pass through specific
anatomical areas. These regions can be used in coordination
with a simple and flexible query language which allows for
arbitrary combinations of these queries using Boolean logic
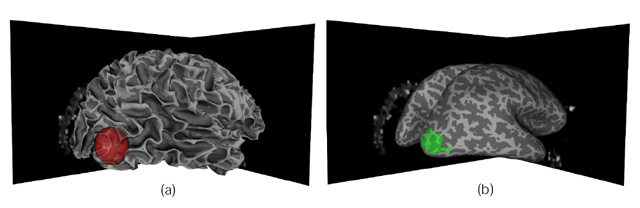
operators. A representation of the cortical surface is provided
for specifying queries of pathways that may be relevant to gray
matter structures, and for displaying activation information
obtained from functional magnetic resonance imaging. By
precomputing the pathways and their statistical properties, we
obtain the speed necessary for interactive question-and-answer
sessions with brain researchers. We survey some questions that
researchers have been asking about tractography data and show
how our system can be used to answer these questions
efficiently.
David Akers | Last updated 14 Jan 2005