|
Neural Process Reconstruction from Sparse User Scribbles |
|
Markus Unger2 |
|
1Harvard University |
2Graz University of Technology |
|
Medical Image Computing and Computer Assisted Intervention (MICCAI) 2011 |
|
|
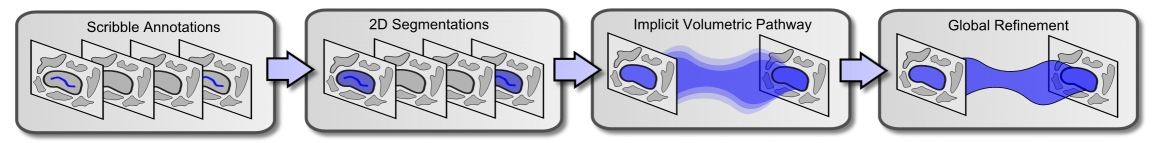
Figure 1: Overview of our method. We assume that we are given scribble annotations indicating a neural process of interest on the first and last slices of an image stack (left). We compute 2D segmentations that contain the scribble annotations and align with strong image edges; these 2D segmentations define hard constraints on our 3D segmentation (middle left). We propagate the 2D segmentations through the image stack according to an implicitly represented volumetric pathway, which we compute based on the dense optical flow between image slices; the interior level sets of this volumetric pathway define soft constraints on our 3D segmentation (middle right). We compute the final 3D segmentation by globally refining the volumetric pathway according to an anisotropic variational segmentation model that aligns with strong in-plane image edges and enforces 3D smoothness (right). |

|
|
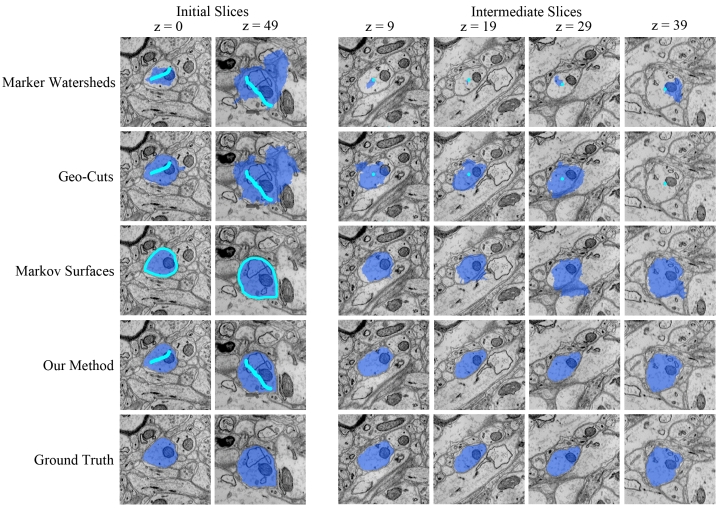
Figure 2: Segmentation results from our method, Markov Surfaces, Geo-Cuts, and Marker-Controlled Watersheds on various slices of a 1024x1024x50 mouse hippocampus EM image stack. Bright blue regions indicate user-provided annotations used to initialize the algorithm, dark blue regions indicate the resulting segmentations. On average, our method is 68% more accurate than previous methods. |
Abstract: We present a novel semi-automatic method for segmenting neural processes in large, highly anisotropic EM (electron microscopy) image stacks. Our method takes advantage of sparse scribble annotations provided by the user to guide a 3D variational segmentation model, thereby allowing our method to globally optimally enforce 3D geometric constraints on the segmentation. Moreover, we leverage a novel algorithm for propagating segmentation constraints through the image stack via optimal volumetric pathways, thereby allowing our method to compute highly accurate 3D segmentations from very sparse user input. We evaluate our method by reconstructing 16 neural processes in a 1024x1024x50 nanometer-scale EM image stack of a mouse hippocampus. We demonstrate that, on average, our method is 68% more accurate than previous state-of-the-art semi-automatic methods.
@inproceedings{roberts:2011,
author = {Mike Roberts AND Won-Ki Jeong AND Amelio Vazquez-Reina AND Markus Unger AND
Horst Bischof AND Jeff Lichtman AND Hanspeter Pfister},
title = {Neural Process Reconstruction from Sparse User Scribbles},
booktitle = {Medical Image Computing and Computer Assisted Intervention (MICCAI '11)},
year = {2011}
}
Acknowledgements: This work was partially supported by the Intel Science and Technology Center for Visual Computing, NVIDIA, the National Institute of Health under Grant No. 1P30NS062685-01 and R01 NS020364-23, the Gatsby Charitable Foundation under Grant GAT3036-Connectomic Consortium, and the National Science Foundation under Grant No. PHY-0835713. We also wish to thank Yongsheng Pan for providing us with the Markov Surfaces source code, and Manhee Lee for helping us to collect ground truth data.
